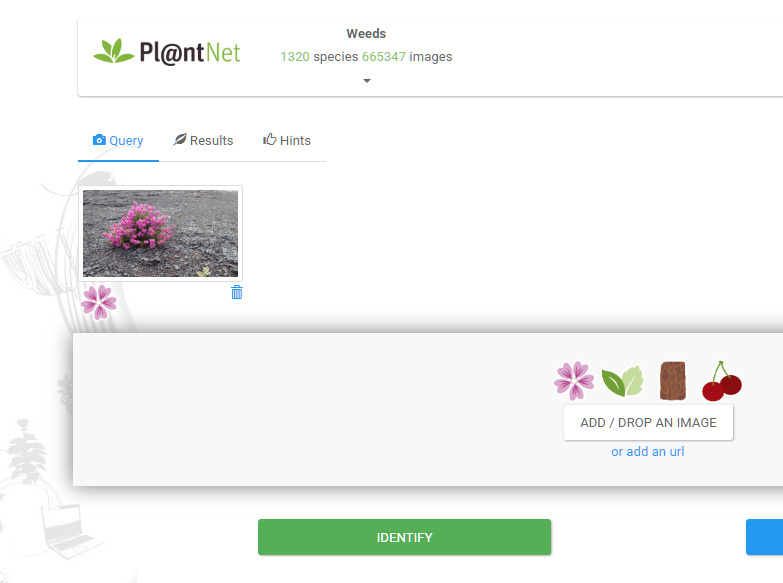
I came across https://identify.plantnet.org the other day and have been quite impressed. In fact, I wish there were similar apps for other species groups. I’ve been using it as the on-line app and my main problem is that it’s a bit slow.
Perhaps a close-up of the flower might be a fairer test?
I note that the community you are using has only 1000+ species; from memory the western european community uses 7000+ species.
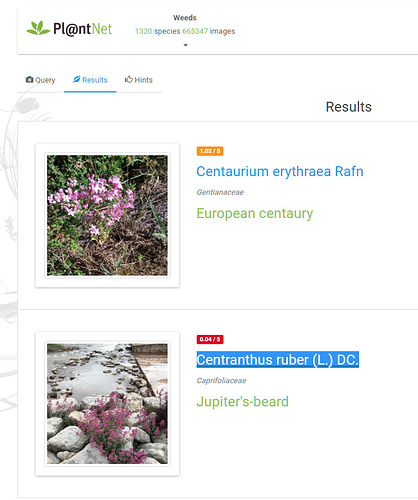
Yes, I tested it as a Weed!.. I also tested a clear pic of a D-b Scabious and it did VERY well choosing D-b S first and Field Sc. second.
You should register in iNat because their AI system is pretty good, from Elephants to Galls. It is hit and miss, generally hit though.
In signing up you do not have to make Observations - just use the ID Software. It just means pretending to Add an Observation. In any case iNat is pretty good. just DON’T leave iSpot!
https://www.inaturalist.org/observations
I took some photographs at Brown Moss in Shropshire earlier this month, so trying them on the Western Europe dataset.
Rumex acetosella - correctly identified
Rumex maritimus - correctly identified.
unidentified thread-leaved plant, probably a capillary-leaved crowfoot, but I thought that these were subaqueous - misidentified as Myriophyllum aquaticum, but the second alternative of Ranunculus trichophyllum could well be correct.
suspected Persicaria mitis- identified with high confidence as Persicaria maculosa (wrong), with Persicaria hydropiper (also wrong - I used the taste test for confirmation) in second place. Persicaria minor, which I haven’t excluded yet, came in 5th. (At least it didn’t focus on the Crassula helmsii dominating the photographs.)
dwarf St. John’s Wort, perhaps depauperate Hypericum perforatum - offered Hypericum tetrapterum, Hypericum humifusum and Hypericum perforatum, in that order.
suspected Veronica scutellata, but I have to check the books to make sure it’s not V. catenata or V. anagallis-aquatica - it agrees with me
Lythrum portula - misidentified as Portulaca oleracea, with the correct identification as the second choice.
laminar-leaved crowfoot - identified as Ranunculus peltatus, so it at least it’s in the right area.
and from Prees Heath
Trifolium arvense - correctly identified
Clinopodium vulgare - it agrees with me
tatty likely Hirschfeldia incana - got it right, but confused me by giving the botanical name Erucastrum incanum and the vernacular name Greek mustard.
Edit: after further thought Ranunculus trichophyllum is was, and the next one was Persicaria minor. And Veronica scutellata and Ranunculus peltatus also seem to be correct.
That is an excellent result, thank you.
Discourse won’t let me edit in more trials. Only the first paragraph comes up on the edit screen.
A good trial - thanks - and pretty good results. These systems are already remarkable and will, I’m sure, only get better: where, then, systems like iSpot which are firmly based on human expertise?
As a matter of interest, did you use the on-line version or one of the mobile apps? How did you find the speed?
Online version. It’s faster than iSpot.
Yes that’s happened a lot recently, I thought it was my browser (and I was the only one). Edit screen is blank and malperfoming - preview is complete.
oh dear…ISpotter’s brains would not have these sorts of failures…
These AI systems are only as good as the identifications that humans make of the images that go into them. Plantnet uses a particularly good system to get good images of various parts of correctly identified plants, the system Google uses does not and instead just tries to use brute force. Just having a huge database of images is no good if they are not correctly identified, indeed it can give very high confidence of wrong ID if most people get the ID wrong in their photos.
There are also issues where there are no good photos of the species in question, a photo I took at the weekend was of a tree growing outside in UK but where there are very likely no photos (yet) on the internet. I asked people at work to identifiy it and of course they tried searching the web with all sorts of tools and came up with nothing even though I gave hints about its native region.
As ever with computers, it’s a matter of “garbage in, garbage out”. Additionally, we are seeing these systems in their infancy and we can expect them to get better as the algorithms for identification improve and as the databases of good quality, well identified photos grow. At the same time systems will poorer algorithms and poorer databases will tend to die out.
I think Pl@ntnet are to be congratulated for creating the current system and for making it freely available. I look forward to seeing it improve with use and, as I noted above, to seeing similar systems becoming available for other species groups.
For the moment, I’ll continue to use it and to add my own expertise - such as it may be - to look critically at the results it offers on the assumption that the photos I am asking it to identify probably don’t show clearly all the features necessary for positive identification: if my photos are garbage …!
Here’s the fault displayed. I don’t think very many people edit comments here (number of Edits is displayed). But for us who do, (for me) it is becoming a nuisance (but please don’t laugh)
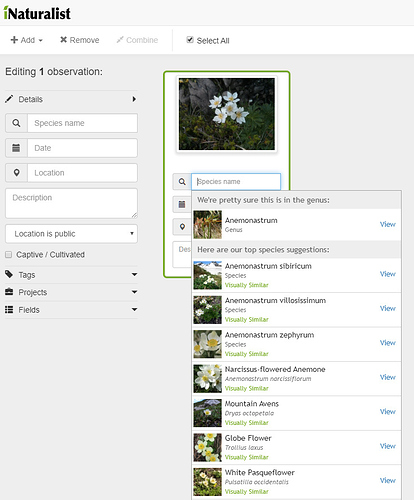
I tried Pla@ntNet for Anemone narcissiflora, my recent post on iSpot.
It offered several white Anemones, the first being A. narcissiflora.
I liked the display of photos already on their database, which was easy to scan through.
As others have said, the limiting feature will be whether of the photo submitted has the relevant features. I am pleased with the result as I don’t take many good photos and mine was clearly good enough.
Next I might try an iSpot post which is so far U/I and see what it makes of it.
Following up … I’ve looked some more at Pl@ntNet. I wonder who, if anyone, is checking the entries.
For example, I found, under Anemone baldensis, some photos of yellow flowers. There were referenced to
Tela Botanica − Yoan MARTIN (cc-by-sa) - Jul 19, 2014.
Clearly they are not A. baldensis (which is white)
I then checked Telabotanica and these photos by Yoan Martin are no longer on the entry for A. baldensis. So, someone is checking that.
Yoan Martin is a contributing botanist with Telabotanica https://www.tela-botanica.org/2018/07/portrait-de-telabotaniste-yoan-martin/
So, as ever, the web is a good resource, but only if we remain alert and critical (in its academic sense).
A cautionary tale
Useful stuff Jo. I was going to suggest that iNat is guilty of allowing such things because of their Research Grade ‘attitude’. But I looked and they are all white. I do think we should use iNat occasionally.
https://www.inaturalist.org/observations?taxon_id=485457
I took the liberty of passing your A.narcissiflora though the iNat AI. It is an excellent photo
Absolutely strongly FIRST in https://identify.plantnet.org/
Here’s iNat
The new user kaldy.xeni might be using identification apps.
Yes, I think you’re right (well spotted) 100% with two of her close ups.
https://www.ispotnature.org/communities/uk-and-ireland/view/observation/790950/
That’s what I thought too -
Unfortunately ID apps don’t always get it right and create a false sense of certainty…I also wonder whether they teach the identifier anything, unlike keying something out, which teaches you the important features to look for next time.